System Biosciences
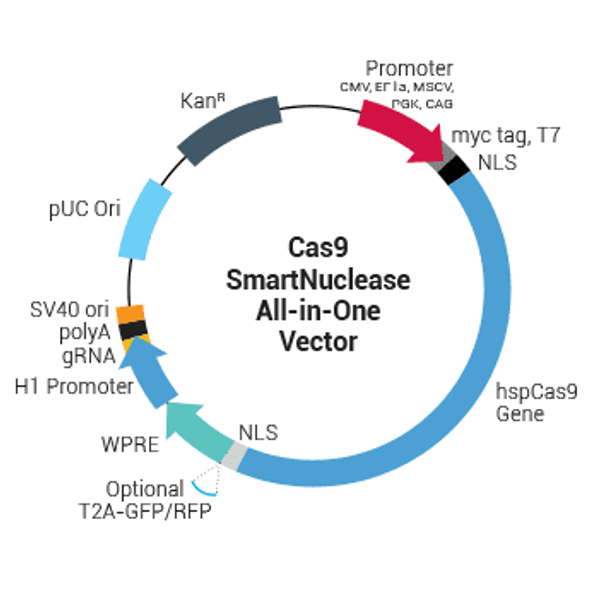
Transfection-ready hspCas9-nickase-T2A-GFP SmartNickase mRNA (Cas9 Nickase mutant with GFP marker)
- SKU:
- CAS534G-1
- Availability:
- Usually Shipped in 5 Working Days
- Size:
- 10 ug
- Shipping Temperature:
- Dry Ice
Description
Transfection-ready hspCas9-nickase-T2A-GFP SmartNickase mRNA (Cas9 Nickase mutant with GFP marker). Cat# CAS534G. Supplier: SBI System Biosciences
Overview
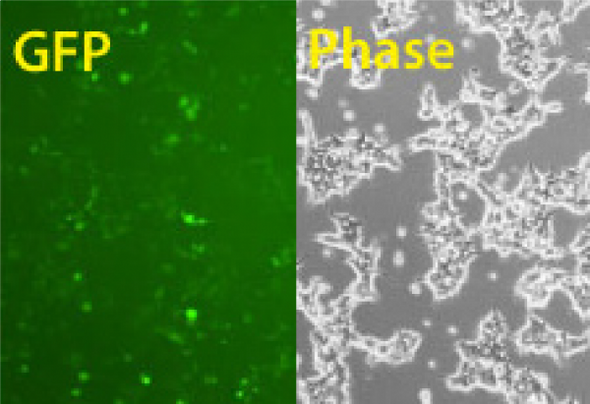
When you want a quick way to verify the efficiency of your Cas9 SmartNickase™ mRNA transfections, SBI offers the hspCas9-T2A-RFP SmartNickase mRNA (a GFP version is also available). This construct delivers coordinated translation of Cas9 and RFP from the same mRNA, enabling the use of the RFP signal to assess transfection efficiency. The nickase mutant of Cas9 is great for pre-clinical and other applications where you need to minimize off-target Cas9 activity.
Unlike the wildtype Cas9 protein which introduces double-strand breaks (DSBs), the Cas9 SmartNickase introduces paired nicks at the gRNA-directed site. Creating nicks favors the higher-fidelity homologous recombination process over non-homologous end joining (NHEJ), with paired nicking shown to reduce off-target activity by 50- to 1,500-fold in cell lines, and to facilitate gene knockout in mice without losing on-target cleavage efficiency1.
As with all of our Cas9 delivery options, the hspCas9-T2A-RFP SmartNickase mRNA is functionally validated and comes backed by our expert technical support team—if you’ve got a genome engineering question just ask by emailing tech@systembio.com.- Ready to use for in vivo genome editing applications
- Enables assessment of Cas9 mRNA transfection efficiency through the RFP signal
- Functionally validated Cas9 SmartNickase
- Backed by expert, easy-to-reach technical support
Even though gene knock-outs can result from DSBs caused by Cas9 alone, SBI recommends the use of HR targeting vectors (also called HR donor vectors) for more efficient and precise mutation. HR donors can supply elements for positive or negative selection ensuring easier identification of successful mutation events. In addition, HR donors can include up to 6-8 kb of open reading frame for gene knock-ins or tagging, and, when small mutations are included in either 5’ or 3’ homology arms, can make specific, targeted gene edits.
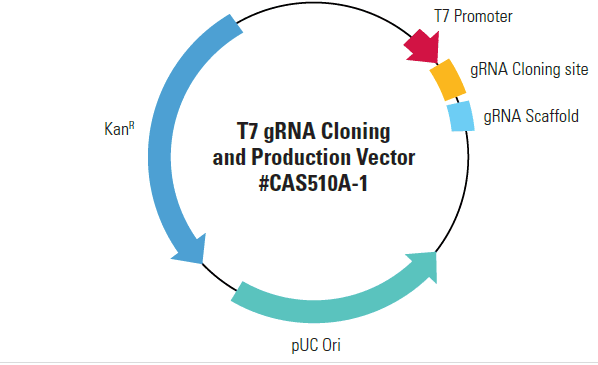
Efficiently generate gRNA with the T7 gRNA SmartNickase Synthesis KitPair your hspCas9-T2A-RFP SmartNickase mRNA with our T7 gRNA SmartNuclease Synthesis Kit, which comes with a pre-linearized, ready-for-cloning gRNA expression vector and all the reagents you need for T7-driven in vitro transcription of your cloned gRNA.
Not sure whether you need a CRISPR/Cas9 plasmid, purified protein, or mRNA?Use this table to choose the CRISPR/Cas9 product that’s right for you:

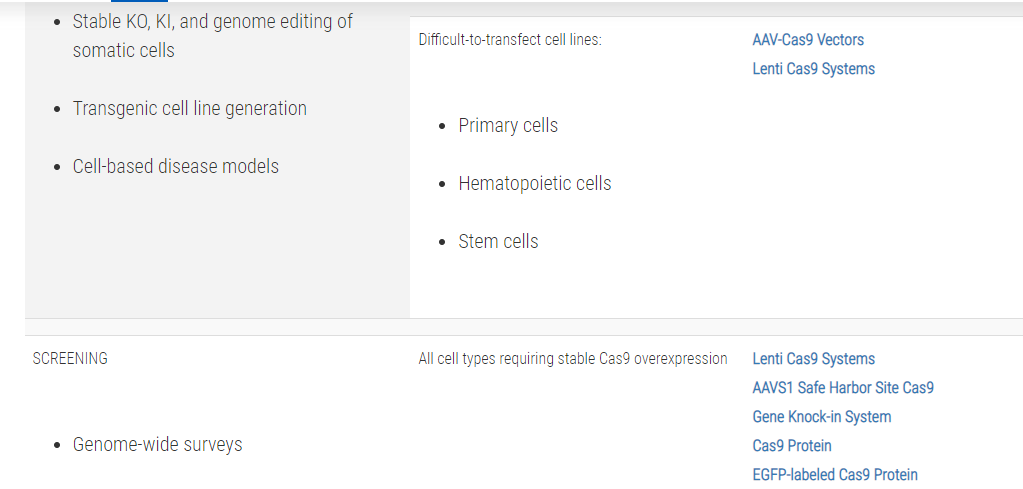
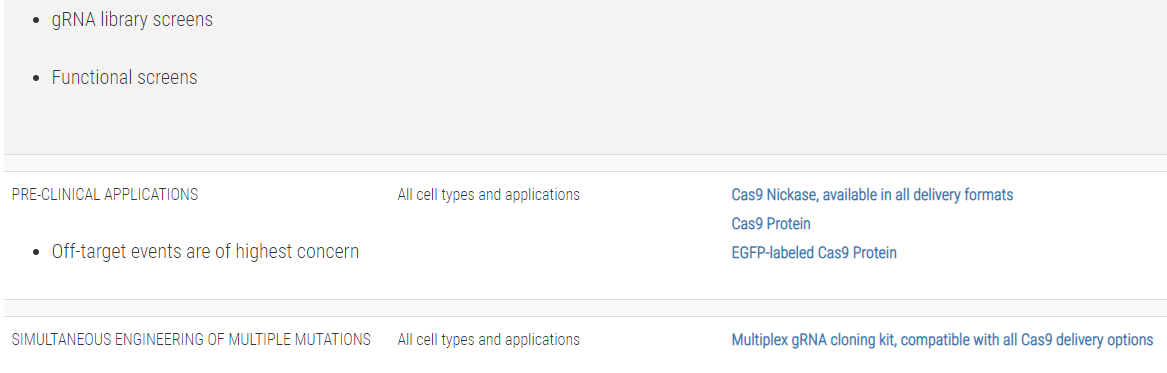
References
- Ran, FA. Genome engineering using the CRISPR-Cas9 system. Nat Protoc. 2013 Oct 24; 8:2281-2308. PMCID: PMC3969860.
How It Works
Genome engineering with CRISPR/Cas9
For general guidance on using CRISPR/Cas9 technology for genome engineering, take a look at our CRISPR/Cas9 tutorials as well as the following application notes:
CRISPR/Cas9 Gene Knock-Out Application Note (PDF) »
CRISPR/Cas9 Gene Editing Application Note (PDF) »
CRISPR/Cas9 Gene Tagging Application Note (PDF) »
CRISPR/Cas9 Basics
Through careful selection of the target sequence and design of a donor plasmid for homologous
recombination, you can achieve efficient and highly targeted genomic modification with CRISPR/Cas9.
The system
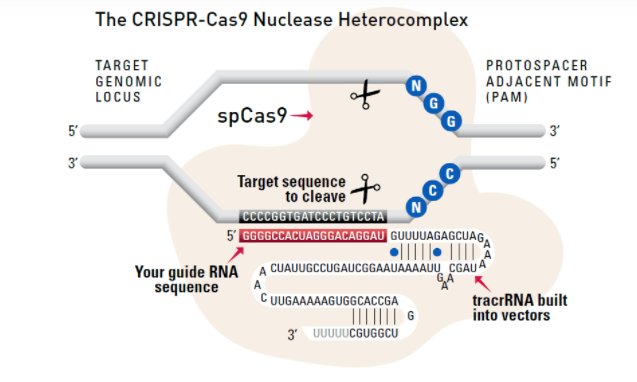
Cas9 protein—uses guide RNA (gRNA) to direct site-specific, double-strand DNA cleavage adjacent to a protospacer adapter motif (PAM) in the target DNA.
gRNA—RNA sequence that guides Cas9 to cleave a homologous region in the target genome. Efficient cleavage only where the gRNA homology is adjacent to a PAM.
PAM—protospacer adapter motif, NGG, is a target DNA sequence that spCas9 will cut upstream from if directed to by the gRNA.
The workflow at-a-glance
DESIGN: Select gRNA and HR donor plasmids. Choice of gRNA site and design of donor
plasmid determines whether the homologous recombination event results in a knock-out,
knock-in, edit, or tagging.
CONSTRUCT: Clone gRNA into all-in-one Cas9 vector. Clone 5’ and 3’ homology arms into HR
donor plasmid. If creating a knock-in, clone desired gene into HR donor.
CO-TRANSFECT or CO-INJECT: Introduce Cas9, gRNA, and HR Donors into the target cells
using co-transfection for plasmids, co-transduction for lentivirus, or co-injection for mRNAs.
SELECT/SCREEN: Select or screen for mutants and verify.
VALIDATE: Genotype or sequence putative mutants to verify single or biallelic conversion.
Supporting Data
Knock-in of GFP at the AAVS1 Safe Harbor Site using Cas9 SmartNickase mRNA
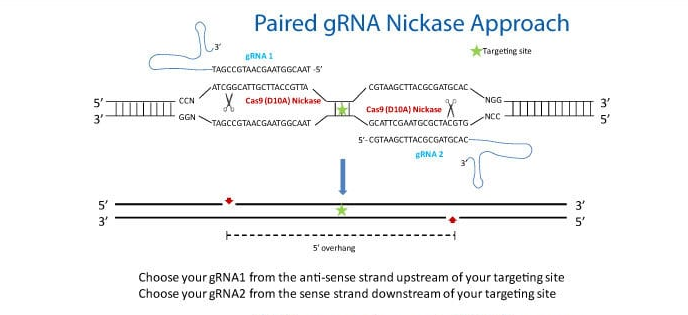

Figure 1. Cas9 SmartNickase mRNA-mediated knock-in of GFP to the AAVS1 Safe Harbor Site. In order to enhance genome editing specificity, paired gRNA with the Cas9 SmartNickase can be used to generate double nicking with 5′ overhang. Please follow the guidelines in the schematic for paired gRNA selection and design—note that only gRNA pairs creating 5′ overhangs with less than 8 bp overlap between the guide sequences were able to mediate detectable indel formation1. To achieve high cleavage efficiency using Cas9 Nickase with paired gRNAs, make sure each gRNA is able to efficiently induce indels when coupled with wide-type Cas9.